Highlighted
All
2024

Fixed-point Encoding and Architecture Exploration for Residue Number Systems
ACM Transactions on Architecture and Code Optimization
·
14 Sep 2024
·
doi:10.1145/3664923
AI-Driven Deep Learning Techniques in Protein Structure Prediction
International Journal of Molecular Sciences
·
01 Aug 2024
·
doi:10.3390/ijms25158426
A Comparative Analysis of SARS-CoV-2 Variants of Concern (VOC) Spike Proteins Interacting with hACE2 Enzyme
International Journal of Molecular Sciences
·
23 Jul 2024
·
doi:10.3390/ijms25158032
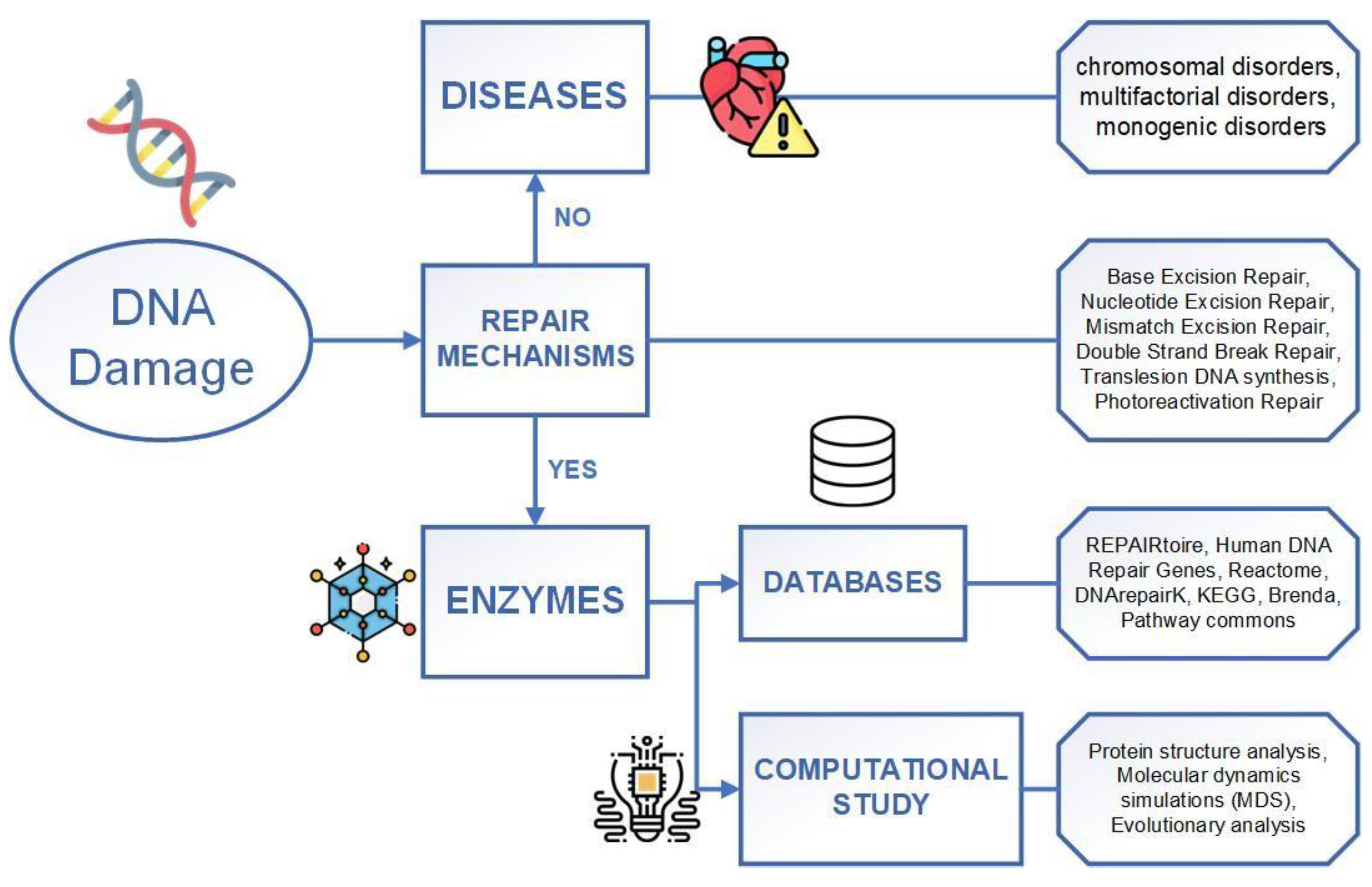
Exploring DNA Damage and Repair Mechanisms: A Review with Computational Insights
BioTech
·
16 Jan 2024
·
doi:10.3390/biotech13010003
2023
Molecular Dynamics Simulations on Coronavirus Variants of Concern (VOC)
The 24th Annual Conference on Information Technology Education
·
11 Oct 2023
·
doi:10.1145/3585059.3611442

JAK3 Y841 Autophosphorylation Is Critical for STAT5B Activation, Kinase Domain Stability and Dimer Formation
International Journal of Molecular Sciences
·
25 Jul 2023
·
doi:10.3390/ijms241511928

Phosphorylation of Tyrosine 841 Plays a Significant Role in JAK3 Activation
Life
·
10 Apr 2023
·
doi:10.3390/life13040981
2022
DG-Labeler and DGL-MOTS dataset: Boost the autonomous driving perception
arXiv
·
07 Jun 2022
·
eid:2-s2.0-85118005743

Computational Study on the Electrostatic Interactions between Uracil‐DNA Glycosylase (UDG) and DNA
The FASEB Journal
·
01 May 2022
·
doi:10.1096/FASEBJ.2022.36.S1.0R568

Computational Study on E-Hooks of Tubulins in the Binding Process with Kinesin
International Journal of Molecular Sciences
·
12 Feb 2022
·
doi:10.3390/ijms23042035

The pH Effects on SARS-CoV and SARS-CoV-2 Spike Proteins in the Process of Binding to hACE2
Pathogens
·
11 Feb 2022
·
doi:10.3390/pathogens11020238
2021

Computational Saturation Mutagenesis of SARS-CoV-1 Spike Glycoprotein: Stability, Binding Affinity, and Comparison With SARS-CoV-2
Frontiers in Molecular Biosciences
·
09 Dec 2021
·
doi:10.3389/fmolb.2021.784303

Computational Study on DNA Repair: The Roles of Electrostatic Interactions Between Uracil-DNA Glycosylase (UDG) and DNA
Frontiers in Molecular Biosciences
·
06 Aug 2021
·
doi:10.3389/fmolb.2021.718587

Computational saturation mutagenesis of SARS-CoV-1 spike glycoprotein: stability, binding affinity, and comparison with SARS-CoV-2
Cold Spring Harbor Laboratory
·
30 Jun 2021
·
doi:10.1101/2021.06.30.450547

The Electrostatic Features of Dengue Virus Capsid Assembly
Journal of Computational Biophysics and Chemistry
·
01 Mar 2021
·
doi:10.1142/S2737416520420089

StructureMan: A Structure Manipulation Tool to Study Large Scale Biomolecular Interactions
Frontiers in Molecular Biosciences
·
11 Jan 2021
·
doi:10.3389/fmolb.2020.627087

Hybrid method for representing ions in implicit solvation calculations
Computational and Structural Biotechnology Journal
·
01 Jan 2021
·
doi:10.1016/j.csbj.2021.01.020

Electrostatic features for nucleocapsid proteins of SARS-CoV and SARS-CoV-2
Mathematical Biosciences and Engineering
·
01 Jan 2021
·
doi:10.3934/MBE.2021120
2020

Spike Proteins of SARS-CoV and SARS-CoV-2 Utilize Different Mechanisms to Bind With Human ACE2
Frontiers in Molecular Biosciences
·
09 Dec 2020
·
doi:10.3389/fmolb.2020.591873

Revealing the Mechanism of SARS-CoV-2 Spike Protein Binding With ACE2
Computing in Science & Engineering
·
01 Nov 2020
·
doi:10.1109/MCSE.2020.3015511

Computational analysis of hereditary spastic paraplegia mutations in the kinesin motor domains of KIF1A and KIF5A
Journal of Theoretical and Computational Chemistry
·
05 Aug 2020
·
doi:10.1142/S0219633620410035

A computational model of ESAT-6 complex in membrane
Journal of Theoretical and Computational Chemistry
·
17 Mar 2020
·
doi:10.1142/S0219633620400027
2019

Using computational approaches to study dengue virus capsid assembly
Computational and Mathematical Biophysics
·
01 Jan 2019
·
doi:10.1515/cmb-2019-0005
